Expertise
Tracking down the species and origin of wood with genetic methods
Hilke Schröder und Céline Blanc-Jolivet | 07.09.2022
When it comes to distinguishing between anatomically similar wood species or making statements about the origin of certain woods, genetic studies are the method of choice. This requires reference samples of woods whose species and origin have been clarified beyond doubt.
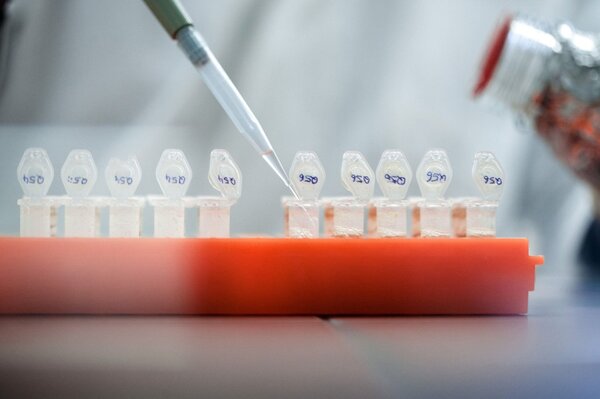
From March 2013, companies importing timber or timber products into the EU must ensure that the products are properly declared, do not come from protected species and do not originate from illegal logging. This is provided for in the European Timber Trade Regulation (EUTR). Among other things, companies must provide information on the country of origin of timber products. But can the traceability documents be trusted? Researchers at the Thünen Institute regularly check timber products using genetic methods and find that in around 20 % of cases the stated region of origin is incorrect.
In order to have suitable comparison material for the genetic examinations, fresh material from more than 50,000 trees from all over the world was collected and analysed in the laboratory.
A long and careful work
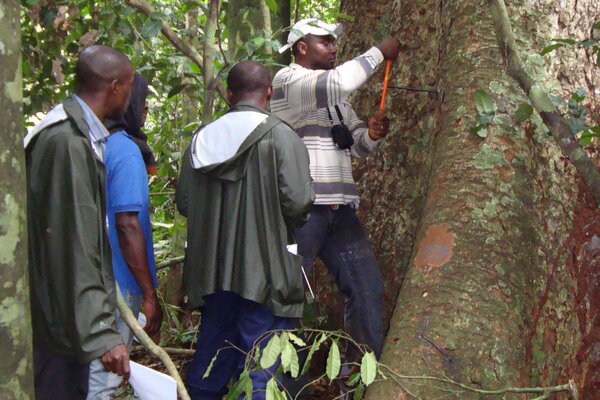
The development of reference data is the result of numerous international collaborations with scientific working groups from wood-producing countries. Trained personnel capable of identifying the respective wood species visit the natural forests with a GPS and sampling tools to collect fresh samples and georeference them. Dried tissue material, or in some cases DNA, is delivered to the respective laboratories where the genetic analyses are carried out.
As in human forensics, DNA analysis can identify a single tree. Modern sequencing technologies provide powerful tools for characterising the genetic patterns of woody species. The vast amount of information from sequencing data can be analysed in such a way that for each wood species only a handful of molecular markers are needed to characterise its geographical genetic pattern. In this way, genetic maps are created for each wood species of interest.
Thünen Institute active in international networks
The Thünen Institute of Forest Genetics is one of the world's leading institutions in genetic wood identification and is active in specialised networks. The exchange of expertise and the identification of the most important species and geographical regions susceptible to illegal logging and timber trade are key to the targeted development of identification methods. Building up reference data is a long-term task that must take into account the state of the timber trade and environmental crime. The database of the Thünen Institute of Forest Genetics now comprises over 54,000 individual samples of 235 tree species from 70 genera. The Institute's international networking with research institutes, particularly in Europe, Africa and South America, has contributed significantly to this, as has its participation in the Global Timber Tracking Network (GTTN).
Distinguishing oak wood from beech wood is reliable and also relatively easy with a magnifying glass and microscope. However, the wood of the different oak species that exist in the world is very similar. This becomes a problem in international trade, because there are rare species that are protected, while other species are traded on a large scale. The Mongolian oak (Quercus mongolica), for example, has a CITES protected status because it forms valuable ecosystems. But even species that are not protected, such as the English oak (Quercus robur), may only be logged in a regulated manner, because in Europe, for example, it is a component of very old natural forests.
In addition to information on species identification, information on the specific origin can also be important. This is the case when trees may be felled in certain regions or countries, but are protected in other countries. An example is the American mahogany (Swietenia macrophylla), a valuable precious wood that occurs in Central and South American natural forests. The tree species is subject to the Washington Convention on International Trade in Endangered Species of Wild Fauna and Flora (CITES) and may currently not be imported into the EU from Bolivia and Belize. Imports from Mexico and Guatemala, on the other hand, are permitted with the appropriate documents because sustainable management is still possible there.
Here, too, genetics is the method of choice to clarify the origin. Even if it is the same species, genetic differences have developed in the individual origins, as there is practically no exchange between the populations due to the distance. However, reference samples from the respective regions are also necessary here in order to find molecular markers in the wood DNA that are typical for the regions.
Wood is dead tissue; its DNA is subject to a constant process of degradation - it degrades. Even the DNA in freshly felled trees is already degraded, but less so than in older or treated wood. In particular, high heat and some chemicals have a strong DNA-degrading effect. This means that only small amounts of DNA can be extracted from wood and wood products - and often only in poor quality. Through our permanent method development, we are nevertheless able to extract and analyse DNA from such highly processed materials as plywood, chipboard, parquet, veneer or even wood pellets.
Molecular markers - the key to success
The genomes of plants and animals are subject to constant change. As a result, the DNA of different species differs in some places. Even within a single species, there are differences in DNA depending on where the individuals come from.
There are two main ways in which DNA varies and these are used to develop molecular markers: Single Nucleotide Polymorphism (SNP) and Length Differences in the DNA Sequence (Insertions or Deletions: InDel).
We make use of these naturally occurring sequence differences between species and also between individuals and specifically search for these variations in the genome. These clearly identifiable, short DNA segments, whose position in the genome we determine via bioinformatic analyses, then serve as molecular markers.



![[Translate to English:] Logo des Bundesministerium für Ernährung und Landwirtschaft](/media/allgemein/logos/BMEL_Logo.svg)