Project
Age determination using Artificial Intelligence
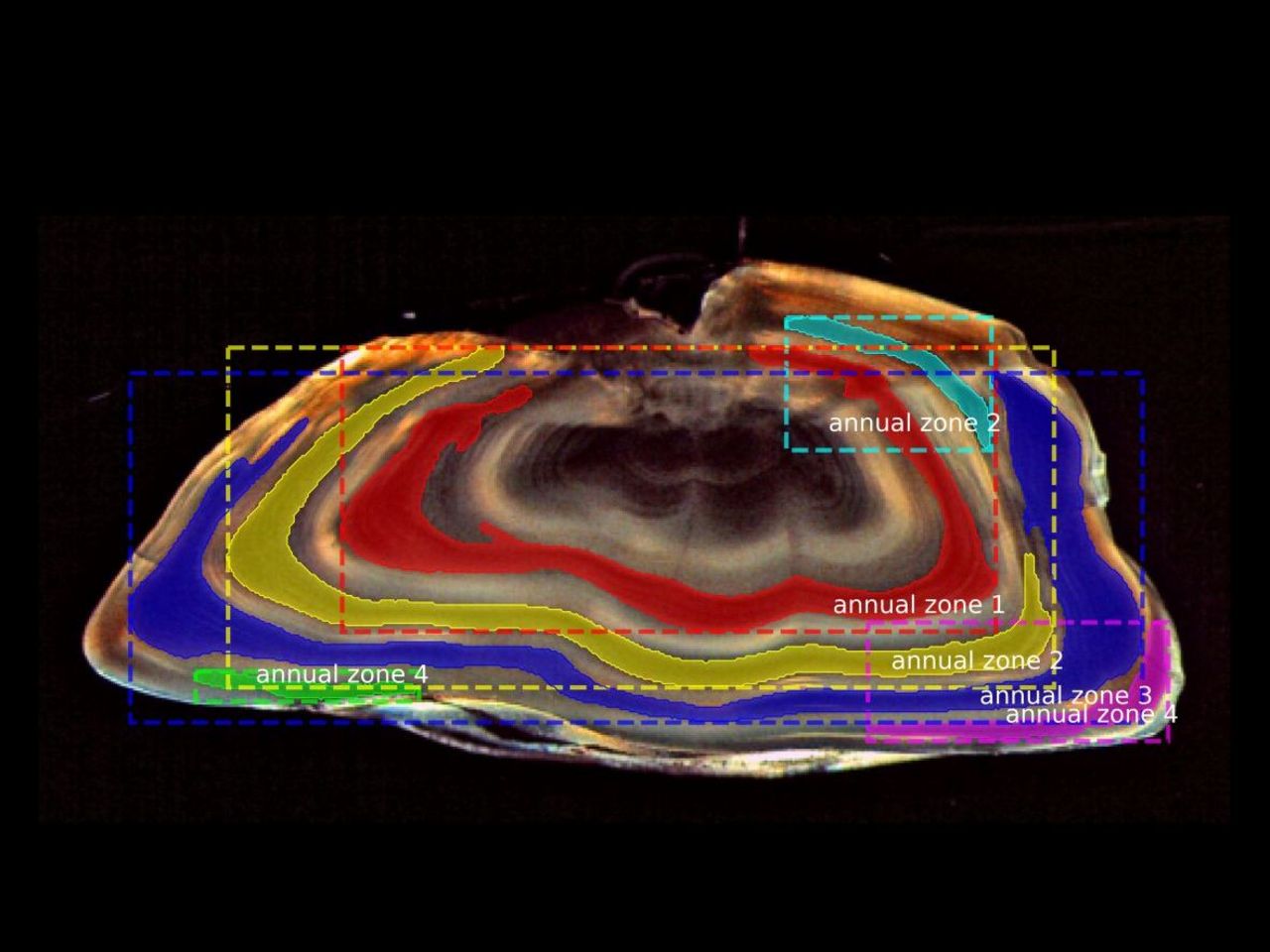
Automated fish age reading and analysis of growth patterns using deep learning
In recent years, the use of artificial intelligence has increased massively in various disciplines. Can this technology also be applied in fisheries science? This question is to be clarified in a PhD project.
Background and Objective
Fish stock assessment relies on accurate estimates of fish age to derive important trends and measurements such as growth and mortality rates. The most widely accepted method has its roots from dendrochronology where the age of a tree is obtained through the counting of rings on its trunk cross-section. In the same manner, fish age can be derived by isolating certain fish structures, particularly the otoliths (ear stones), where growth ring patterns similar to tree-rings are formed at regular intervals in response to changing seasons.
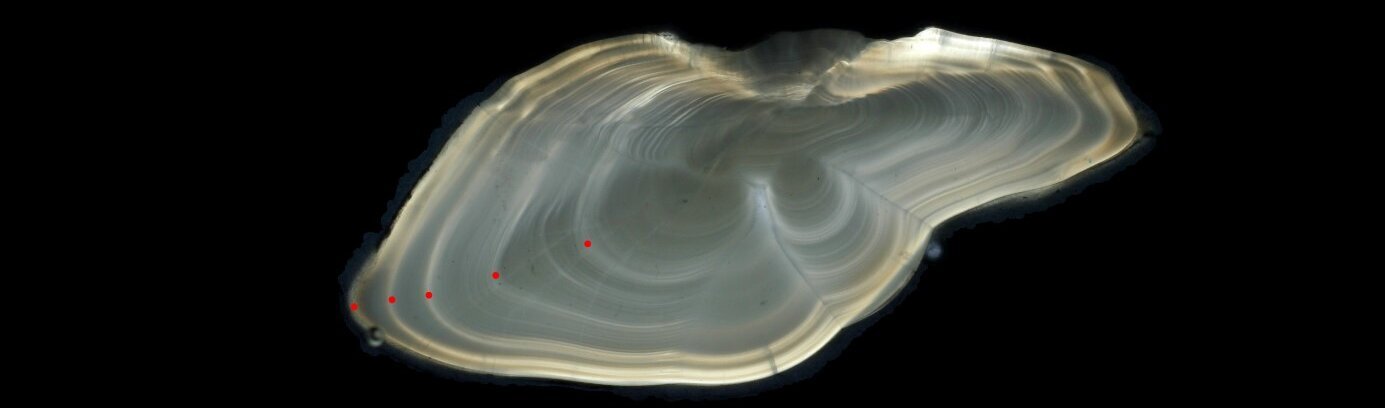
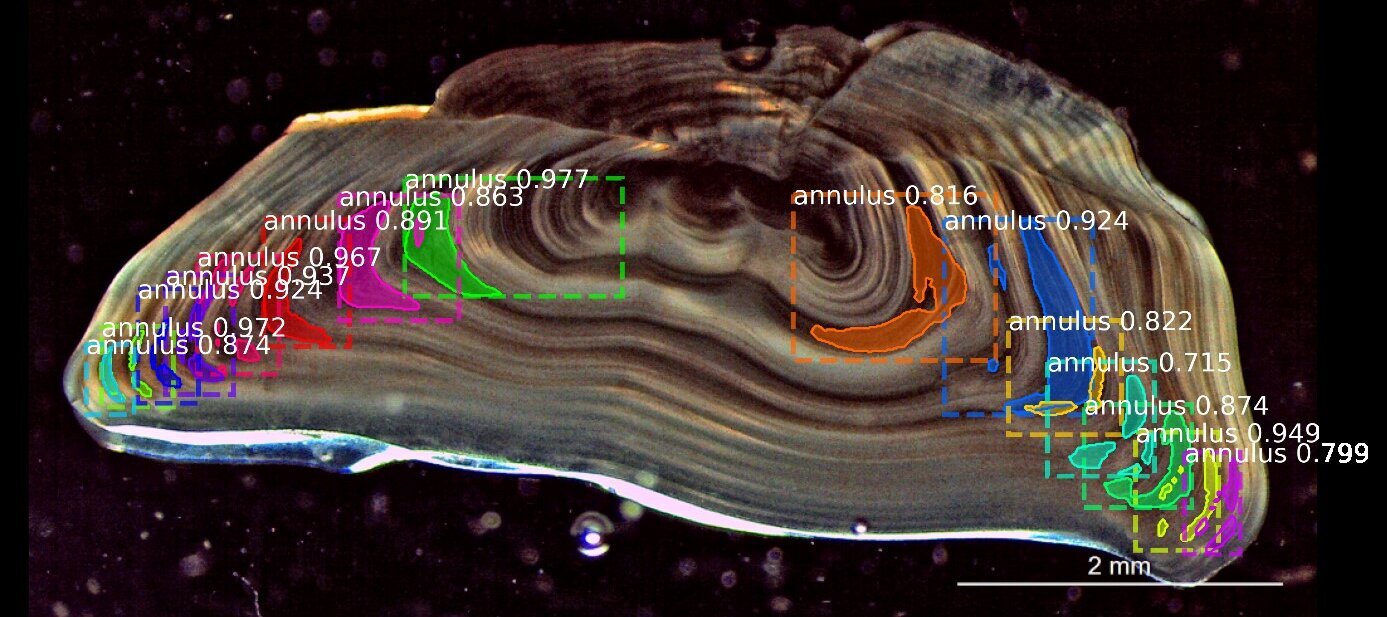
To illustrate the manual age reading process, Figure 1 shows an image of an otolith where the so-called annuli (annual growth rings) were annotated by an expert age reader and the age was estimated to be 5 years. To make the method more accurate and less prone to subjective errors, the age readers undergo years of training and workshops to learn various guidelines on how to correctly distinguish the annual growth rings which can differ from one species (or even stock) to another. The goal is to prevent over- and under-estimation of fish ages since this can cause detrimental effects on fish stock management.
Due to the subjective and logistic limitations of this conventional method, it is therefore timely to utilize the advances in the field of AI and to make use of various deep learning algorithms to provide objective estimates of fish age from otolith images.
Links and Downloads
He K., Gkioxari G., Dollár P. and Girshick R. 2017. Mask R-CNN. IEEE International Conference on Computer Vision (ICCV), Venice, 2017, pp. 2980-2988, doi: 10.1109/ICCV.2017.322.
Ronneberger O., Fischer P., Brox T. 2015. U-Net: Convolutional Networks for Biomedical Image Segmentation. In: Navab N., Hornegger J., Wells W., Frangi A. (eds) Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015. MICCAI 2015. Lecture Notes in Computer Science, vol 9351. Springer, Cham. doi.org/10.1007/978-3-319-24574-4_28
Thünen-Contact

Involved Thünen-Partners
Duration
3.2021 - 5.2024
More Information
Project status:
ongoing
Publications
- 0
Cayetano A, Stransky C, Birk A, Brey T (2024) Fish age reading using deep learning methods for object-detection and segmentation. ICES J Mar Sci: Online First, Feb 2024, DOI:10.1093/icesjms/fsae020

![[Translate to English:] [Translate to English:]](/media/_processed_/7/1/csm_IMG_7977_large_1defaf5de1.jpg)

![[Translate to English:] Logo des Bundesministerium für Ernährung und Landwirtschaft](/media/allgemein/logos/BMEL_Logo.svg)